Browse Articles
Research Article
Namrata Setia, Robert A Goulart, Gladywn Leiman, Christopher N Otis, Rukmini Modem, Liron Pantanowtiz
Research Article
Torill Sauer, Kristin Ebeltoft, Mette Kristin Pedersen, Rolf Kåresen
Editorial
Inderpreet Dhillon, Martha B. Pitman, Richard M. DeMay, Pamela Archuletta, Vinod B. Shidham
Research Article
Elsa Beraki, Torill Sauer
Research Article
Hannah A. Kastenbaum, Walid E. Khalbuss, Raymond E. Felgar, Ronald Stoller, Sara E. Monaco
Case Report
Namrata Setia, Peter Ghobrial, Pantanowitz Liron
Case Report
Chetna N. Purohit, Marilyn M. Bui, Ardeshir Hakam
Case Report
Renuka Gahine, Vijaya Sudarshan, Nighat Hussain, Chandani Krishnani
Research Article
Papa Dasari, S Rajathi, Surendra V Kumar
Research Article
Bharat Rekhi, Dulhan Ajit, Santhosh K Joseph, Sonali Gawas, Kedar K Deodhar
Case Report
Tatiana V. Yakoushina, Ehud Lavi, R. S. Hoda
Case Report
Richard H Siderits, Osman Ouattara, Alan Marcus, Hong Guang Gao, Hong Bing Deng, Janusz Godyn
Case Report
Shaoying Li, Zhijie Yan, Nirag Jhala, Darshana Jhala
Original Article
Samer N. Khader, Kathie Schlesinger, Josh Grossman, Richard I. Henry, Mark Suhrland, Amy S. Fox
Original Article
Torill Sauer
Original Article
Sara E. Monaco*, Matthew J. Schuchert, Walid E. Khalbuss
Original Article
Pam Michelow*, Ingrid Hartman, Doreen Schulze, Stella Lamla-Hillie, Sophie Williams, Simon Levin, Cynthia Firnhaber
Case Report
Liron Pantanowitz*, Michael Kuperman, Robert A. Goulart
Original Article
Prabodh K Gupta
Case Report
Hatem Q. Al-Maghraby, Walid E. Khalbuss, Uma N. M. Rao, Kathleen Cieply, Sanja Dacic, Sara E. Monaco
Case Report
Ruchika Gupta, Sandeep R. Mathur, Venkateswaran K. Iyer, Sudheer Kumar A, Amlesh Seth
Review
G. Denice Smith, Matt Riding, Kim Oswald, Joel S. Bentz
Original Article
J Rimiene, J Petronytė, Z Gudleviciene, Smailytė Giedrė, Krasauskaite Ingrida, A Laurinavicius
Original Article
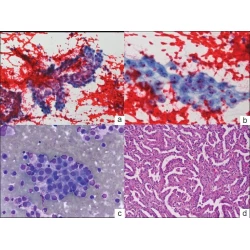
Vinod B Shidham, George Varsegi, Krista D’Amore













Commentary
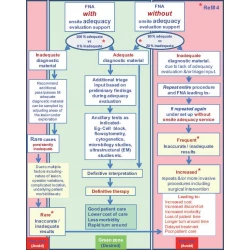
Adequate reimbursement is crucial to support cost-effective rapid on-site cytopathology evaluations
Mousa A. Al-Abbadi, Leonard I. Bloom, Lisa A. Fatheree, Lori A. Haack, Gerald Minkowitz, David C. Wilbur, Marshall R. Austin
Full text |
|  PDF
PDF 